Co-Infection Technical Bulletin
Contents
The Importance of Co-Infections
As we focus on diagnosing and stopping the spread of COVID-19 during this global pandemic, it’s important to remember that co-infections with other respiratory pathogens may have an important role to play in determining the risk of more severe disease and in guiding management strategies for physicians. An epidemiologic study found 11% of viral respiratory infections in a group of urban in- and outpatients involved co-infection with another virus, and the incidence of co-infection (18%) was even more common in children ≤ 5 years old.14 During both pandemic and seasonal influenza episodes, bacterial and viral co-infections led to more severe clinical outcomes and death, especially among the elderly and young children.11 In the intensive care (ICU) setting, the clinical course for patients with community acquired pneumonia (CAP) and viral-bacterial co-infections is far more complicated than that of patients with a single pathogen.17
Co-Infections and COVID-19
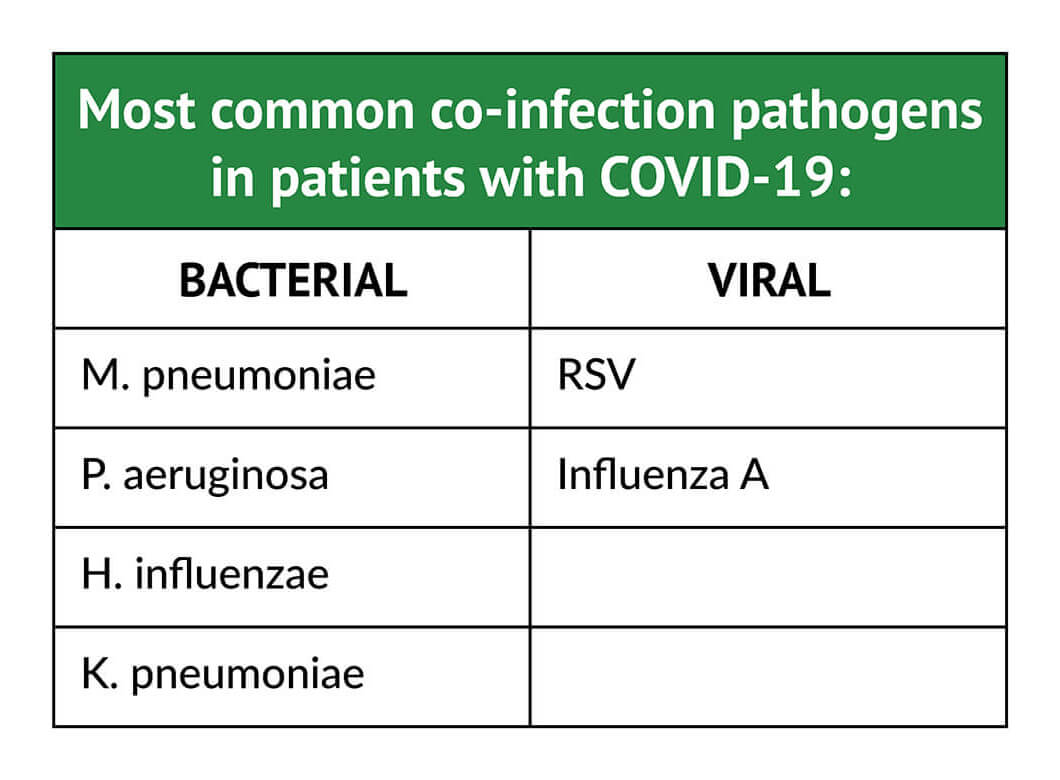
New information is now emerging about co-infections in people infected with COVID-19. A recently published systematic review of 30 studies that included 3,834 people diagnosed with COVID-19 found that 7% had laboratory-confirmed bacterial co-infections, increasing to 14% in studies that included only ICU patients.13 Furthermore, the co-infecting pathogens in patients with COVID-19 are different from those identified in patients during the 2009 (H1N1) influenza pandemic.2,13 In patients with COVID-19, the most common co-infecting bacterial pathogen was Mycoplasma pneumoniae (M.pneumoniae), followed by Pseudomonads aeruginosa (P. aeruginosa), Hemophilus influenzae (H.influenzae), and Klebsiella pneumoniae (K.pneumoniae); respiratory syncytial virus (RSV) and influenza A were the most common co-infecting viral pathogens.13 In COVID-positive outpatients and individuals evaluated in emergency departments who were not admitted to the hospital, the incidence of co-infection may be as high as 20% to 26%.12
 There is some evidence that having a co-infection negatively impacts the health outcomes of people infected with COVID-19. Co-infections may increase the likelihood of being admitted to the hospital4 and, in one study, COVID-19 positive patients with co-infections were more likely to die than patients without co-infections.13 Even in individual studies that did not find a correlation between the presence of bacterial, viral or fungal co-infections and more severe disease, death or ICU admissions for COVID-positive patients1,19, identification of the coinfecting pathogens helped to guide diagnoses and treatments for patients19 and stratification by age suggested that pre-school children were potentially at higher risk for poor outcomes.1 More studies are clearly needed to determine the extent to which co-infections impact the clinical presentation and outcomes for COVID-19- positive patients. Nonetheless, experts have recommended that if influenza and respiratory syncytial virus (RSV) are circulating in the community, it is reasonable to also test for these viruses when testing for COVID-19, as this could have management implications.8
There is some evidence that having a co-infection negatively impacts the health outcomes of people infected with COVID-19. Co-infections may increase the likelihood of being admitted to the hospital4 and, in one study, COVID-19 positive patients with co-infections were more likely to die than patients without co-infections.13 Even in individual studies that did not find a correlation between the presence of bacterial, viral or fungal co-infections and more severe disease, death or ICU admissions for COVID-positive patients1,19, identification of the coinfecting pathogens helped to guide diagnoses and treatments for patients19 and stratification by age suggested that pre-school children were potentially at higher risk for poor outcomes.1 More studies are clearly needed to determine the extent to which co-infections impact the clinical presentation and outcomes for COVID-19- positive patients. Nonetheless, experts have recommended that if influenza and respiratory syncytial virus (RSV) are circulating in the community, it is reasonable to also test for these viruses when testing for COVID-19, as this could have management implications.8
PCR Testing
Nucleic acid based tests (NATs) are based on the polymerase chain reaction (PCR) and detect pathogen specific DNA or RNA sequences rather than antigens and antibodies. A large variety of NATs are currently available for diagnostic purposes. PCR is the most well-developed molecular technique to date for detecting a broad spectrum of viral and bacterial pathogens and for profiling antimicrobial resistance, and its use has increased significantly over the past several years (Figure 3A).10, 15
PCR is an enzyme driven process that amplifies short regions of DNA in vitro. When at least a partial sequence of the target DNA is known beforehand, oligonucleotide primers can be synthesized that bind specifically to the target sequences. In PCR, the target DNA is copied by a DNA polymerase enzyme and can then be exponentially amplified through a process involving multiple cycles of heating and cooling in a thermocycler (Figure 4). In reverse transcription PCR (RT-PCR), RNA is reverse transcribed to a single-stranded cDNA, prior to amplification. In quantitative real-time PCR, amplification and detection of amplified products are carried out together in a single reaction vessel. Using different types of fluorescent probes, a fluorescent signal is emitted during each amplification cycle only when target sequences are present. The intensity of the signal increases in proportion to the amount of amplified product.18 In addition to its quantitating ability, real-time PCR is faster and more reproducible than conventional PCR.
Traditional diagnostic techniques used for detecting bacterial and viral pathogens, such as cultures, antigen detection, viral isolation, serologic assays, and direct fluorescent antibody tests are time-consuming and can yield variable results16 that may take days to finalize and report, delaying accurate diagnoses and the implementation of targeted antimicrobial therapies. PCR has a shorter turn-around time and a higher sensitivity for viral pathogens16 and can detect the presence of microbial pathogens far below the level required by other techniques.18 Multiplex PCR, which enables the simultaneous detection of several target sequences by incorporating multiple sets of primers, is being increasingly relied on for the accurate detection of respiratory pathogens (Figure 3B), and several mPCRs have been approved by the US Food and Drug Association (FDA) for that purpose.5
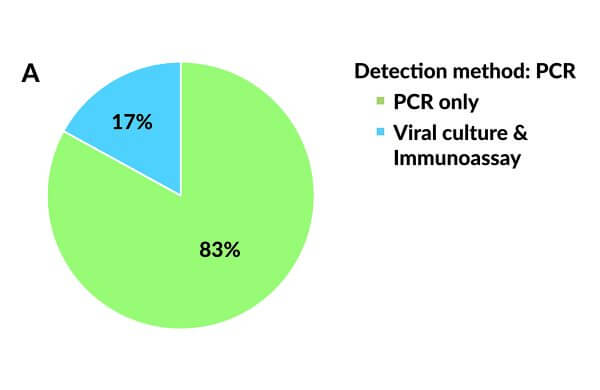
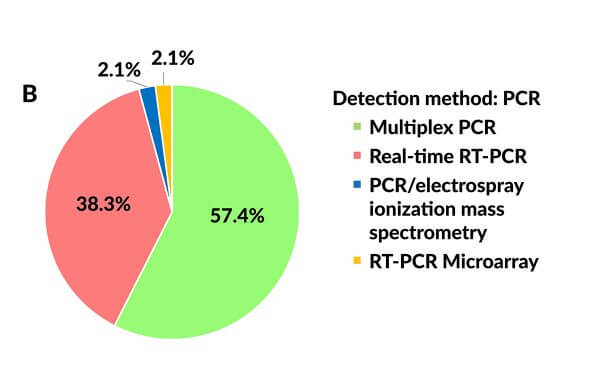
Figure 3. Applied laboratory assays for detecting human metapneumovirus. (A) Pie chart showing the use of viral culture and immunoassays vs reverse transcriptase polymerase chain reaction. (RT-PCR). (B) Pie chart showing the use of different PCR-based assays.
Multiplex PCR and Co-Infections
Multiplex PCR can test for a broader panel of pathogens and co-infections. In a systematic review of 20 studies that included 5,510 patient samples, 30% of which were obtained from children, three mPCRs were found to be highly accurate and provided early identification of respiratory virus pathogens that could be beneficial for management and treatment decisions.9 The combined sensitivity (true positives) of the three mPCRs for detection of influenza A (FluA) and B (FluB), respiratory syncytial virus (RSV), and human metapneumovirus (hMPV) were between 91 and 95%. Most notably for FluA, the combined specificity (true negatives) of 98% was higher than that obtained with the rapid influenza diagnostic test (which can be performed in 15-30 minutes).3 For the clinician, this means greater confidence that a person does not have FluA (if the prevalence of FluA in the community is low). mPCR of a single respiratory sample obtained from hospitalized patients with CAP doubled the detection of bacterial pathogens from 39% to 87% and also allowed for a determination of bacterial load (which could help distinguish between contamination and colonization).7 Furthermore, mPCR detected pathogens in 77.6% of patients who had already received antibiotics compared with a detection rate of only 32.1% by routine culture, providing useful information that might alter a clinician’s treatment strategy.7
Clinical Advantages of mPCR for the Detection of Respiratory Pathogens
When combined with clinical assessment, laboratory and radiographic studies, the high sensitivity and specificity of mPCRs may provide several benefits to the clinician. The rapid results could lead to greater precision in choosing antimicrobial treatments, decreasing the use of empirical antimicrobial and, as a consequence, reducing antimicrobial resistance.9 Both physicians and patients would benefit from less repetitive testing and elimination of wait times for traditional laboratory results.18 Early identification of viral infections may also decrease the use of antibiotics as well as reduce the potential of asymptomatic spreading to vulnerable populations from asymptomatic individuals with viral shedding.6
PCR for the Detection of Other Pathogens
Several PCR-based assays for the detection of specific microbes have gained wide acceptance in clinical practice, including Mycobacterium tuberculosis (M. tuberculosis), Chlamydia trachomatis (C. trachomatis) from both genital and urine samples, herpes simplex virus, cytomegalovirus, hepatitis virus, and HIV to name a few (Table 1).18 A current (2020) list of FDA-approved nucleic acid based tests for the detection of microbial pathogens can be found at: https://www.fda.gov/medical-devices/vitro-diagnostics/nucleic-acid-based-tests.
Assurance Scientific Laboratories Panel for the Detection of Pathogens
At Assurance Scientific Laboratories, in addition to COVID-19 testing, we offer several respiratory panels for the detection of co-infecting viral and bacterial pathogens (COVID respiratory panel lite, COVID respiratory panel, COVID respiratory panel plus). In addition, we offer a wide range of pathogen detection panels for urinary tract and wound infections, sexually transmitted diseases, vaginitis, and gastrointestinal pathogens, several of which include antimicrobial sensitivity testing. Testing for candida and CMV are also available. Stand-alone antimicrobial sensitivity testing for a wide range of antibiotics is also available.
General Guidelines
Laboratory test results should always be considered in the context of clinical observations and epidemiological data (such as local prevalence rates and current outbreak/epicenter locations) in making a final diagnosis and patient management decisions. With any test, the possibility of false positive and false negative results should always be considered, and the impact on patient management decisions and clinical outcomes should be carefully weighed.
References
- Asner, S. A., Science, M. E., Tran, D., Smieja, M., Merglen, A., & Mertz, D. (2014). Clinical disease severity of respiratory viral co-infection versus single viral infection: a systematic review and meta-analysis. PLoS One, 9(6), e99392. doi:10.1371/journal.pone.0099392.
- Blyth, C. C., Webb, S. A., Kok, J., Dwyer, D. E., van Hal, S. J., Foo, H., . . . Investigators, C. M. (2013). The impact of bacterial and viral co-infection in severe influenza. Influenza Other Respir Viruses, 7(2), 168-176. doi:10.1111/j.1750-2659.2012.00360.x.
- Chartrand, C., Leeflang, M. M., Minion, J., Brewer, T., & Pai, M. (2012). Accuracy of rapid influenza diagnostic tests: a meta-analysis. Ann Intern Med, 156(7), 500-511. doi:10.7326/0003-4819-156-7-201204030-00403.
- Davis, B., Rothrock, A. N., Swetland, S., Andris, H., Davis, P., & Rothrock, S. G. (2020). Viral and atypical respiratory co-infections in COVID-19: a systematic review and meta-analysis. J Am Coll Emerg Physicians Open. doi:10.1002/emp2.12128.
- Diaz-Decaro, J. D., Green, N. M., & Godwin, H. A. (2018). Critical evaluation of FDA-approved respiratory multiplex assays for public health surveillance. Expert Rev Mol Diagn, 18(7), 631-643. doi:10.1080/14737159.2018.1487294.
- Esposito, S., Mencacci, A., Cenci, E., Camilloni, B., Silvestri, E., & Principi, N. (2019). Multiplex Platforms for the Identification of Respiratory Pathogens: Are They Useful in Pediatric Clinical Practice? Front Cell Infect Microbiol, 9, 196. doi:10.3389/fcimb.2019.00196.
- Gadsby, N. J., Helgason, K. O., Dickson, E. M., Mills, J. M., Lindsay, D. S., Edwards, G. F., . . . Escmid Study Group for Legionella Infections, B. S. (2016). Molecular diagnosis of Legionella infections–Clinical utility of front-line screening as part of a pneumonia diagnostic algorithm. J Infect, 72(2), 161-170. doi:10.1016/j.jinf.2015.10.011.
- https://www.uptodate.com/contents/coronavirus-disease-2019-covid-19-diagnosis?search=coinfection%20%20and%20covid19&source=search_result&selectedTitle=1~150&usage_type=default&display_rank=1. (2020).
- Huang, H. S., Tsai, C. L., Chang, J., Hsu, T. C., Lin, S., & Lee, C. C. (2018). Multiplex PCR system for the rapid diagnosis of respiratory virus infection: systematic review and meta-analysis. Clin Microbiol Infect, 24(10), 1055-1063. doi:10.1016/j.cmi.2017.11.018.
- Jeong, S., Park, M. J., Song, W., & Kim, H. S. (2020). Advances in laboratory assays for detecting human metapneumovirus. Ann Transl Med, 8(9), 608. doi:10.21037/atm.2019.12.42.
- Joseph, C., Togawa, Y., & Shindo, N. (2013). Bacterial and viral infections associated with influenza. Influenza Other Respir Viruses, 7 Suppl 2, 105-113. doi:10.1111/irv.12089.
- Kim, D., Quinn, J., Pinsky, B., Shah, N. H., & Brown, I. (2020). Rates of Co-infection Between SARS-CoV-2 and Other Respiratory Pathogens. JAMA. doi:10.1001/jama.2020.6266.
- Lansbury, L., Lim, B., Baskaran, V., & Lim, W. S. (2020). Co-infections in people with COVID-19: a systematic review and meta-analysis. J Infect, 81(2), 266-275. doi:10.1016/j.jinf.2020.05.046.
- Nickbakhsh, S., Thorburn, F., B, V. O. N. W., Mc, M. J., Gunson, R. N., & Murcia, P. R. (2016). Extensive multiplex PCR diagnostics reveal new insights into the epidemiology of viral respiratory infections. Epidemiol Infect, 144(10), 2064-2076. doi:10.1017/S0950268816000339.
- Siow, W. T., Koay, E. S., Lee, C. K., Lee, H. K., Ong, V., Ngerng, W. J., . . . Phua, J. (2016). The Use of Polymerase Chain Reaction Amplification for the Detection of Viruses and Bacteria in Severe Community-Acquired Pneumonia. Respiration, 92(5), 286-294. doi:10.1159/000448555.
- Vemula, S. V., Zhao, J., Liu, J., Wang, X., Biswas, S., & Hewlett, I. (2016). Current Approaches for Diagnosis of Influenza Virus Infections in Humans. Viruses, 8(4), 96. doi:10.3390/v8040096.
- Voiriot, G., Visseaux, B., Cohen, J., Nguyen, L. B., Neuville, M., Morbieu, C., . . . Timsit, J. F. (2016). Viral-bacterial coinfection affects the presentation and alters the prognosis of severe community-acquired pneumonia. Crit Care, 20(1), 375. doi:10.1186/s13054-016-1517-9.
- Yang, S., & Rothman, R. E. (2004). PCR-based diagnostics for infectious diseases: uses, limitations, and future applications in acute-care settings. Lancet Infect Dis, 4(6), 337-348. doi:10.1016/S1473-3099(04)01044-8.
- Zhu, X., Ge, Y., Wu, T., Zhao, K., Chen, Y., Wu, B., . . . Cui, L. (2020). Co-infection with respiratory pathogens among COVID-2019 cases. Virus Res, 285, 198005. doi:10.1016/j.virusres.2020.198005.
Ready to take charge of your lab
services?
START YOUR LAB
USE OUR LAB
© Copyright 2024 | All Rights Reserved | Privacy Policy | Terms of Use | Do not sell or share my personal information


